So now comes the challenging bit.
You've gathered all the information you can. You've used some of the services to try and interpret your data. What now?
Let's face the problem head-on. As ME patients, we don't have a lot of focus. It's hard to devote any kind of energy to anything, and that includes intellectual energies. I'm a devoted author who loves to write, but it's become very hard for me to work on anything long-term because juggling that many variables eventually gets to feel like work; it loses its entertainment value and I'm left feeling like writing fantasy is a job I have to push through.
Short articles like this are usually within my grasp. As you'll see by looking at my closing statements, I usually just stop when I get to feeling like that and say, "see you next time!" We can't always do that in other arenas of our lives. I've always had a lot of difficulty stopping a job partway through and picking it up coherently, later, which is why my best stories are always the ones completed within a few days. I believe this is an aspect of my personality rather than something ME has done to my brains.
So when you're handed a list of SNPs in the hundreds of thousands - or at least, when I am - my brain looks at that long list and fizzles out like a candle in a high wind.
So it's tempting to look at the work that someone else has done and say, "well, I guess that seems reasonable." It's also very tempting to simplify and say that any negative SNP means disaster, or that none of it has significant impact at all. If it's some one way and some the other, and sometimes SNP X only has a significant impact if SNP Y is also involved, it gets complicated.
Let's take our good old friend, CBS C699T. After all, I have the dreaded red box for that SNP, so it's in my best interest to find out what kind of impact that will have on my life. I'm going to show you the process I used to determine the meaningfulness of this SNP.
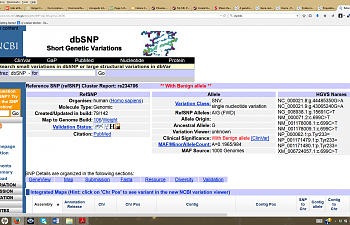
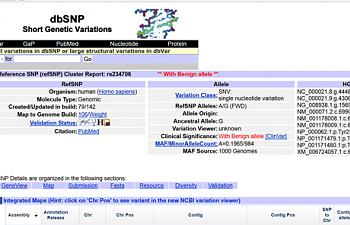
1) Go to pubmed, scroll down the 'Search Entrez' box until it says 'SNP' and enter the SNP ID number of your SNP (that rs#### thingie). When you hit enter, a list of several different links will pop up with this rs#. Click the SNP ID number that says 'Homo sapiens' next to it (the first link.) This will pop up:
NCBI has an enormous amount of information regarding each SNP, but it's a little hard to wade through. We're gonna take this one step at a time, though.
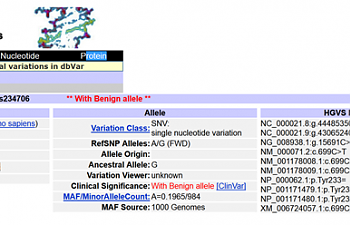
Let's take a closer look at the middle column that says "allele":
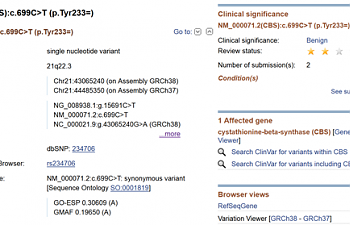
2) If you click where it says 'ClinVar', it will take you here:
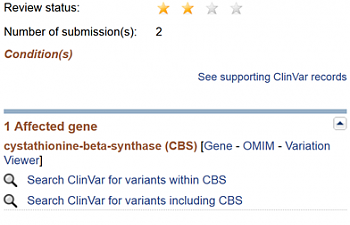
This is more detailed information regarding this SNP. Look on the right side:
It says that the clinical significance is benign, and lists no conditions associated with this SNP. The two stars say that more than one researcher has provided evidence that this SNP is benign. So you don't have to worry about this SNP, at least as far as NCBI is concerned.
Okay. Let's assume we don't trust their bloody medical opinions. Frankly, if you have utmost faith in The Man and the Establishment and you have this illness, you haven't been paying enough attention.
It does say that the cystathionine-beta-synthase gene is affected. Let's look into that a little more.
3) Click where it says 'Gene' next to CBS's name. This will take you to a page devoted to the CBS gene. If you scroll down this new page:
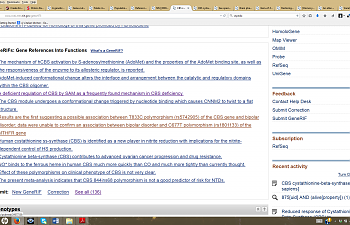
You can find articles about the CBS gene like there's no tomorrow. 136 regarding references into functions as of this posting! That is pretty darned impressive, but not possible to read through. You can, however, open up all of those by going here:
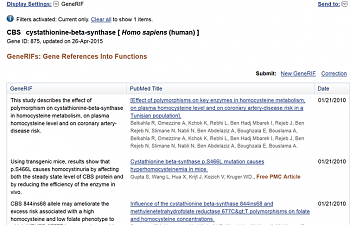
4) Open up 'see all' at the bottom of the page. Then, you won't just see the articles, but summaries of what each of them convey. This makes them very quick to scroll through and see if you can find anything interesting.
Look at those handy little summaries on the left! How lovely of them. <3
5) Determine what looks 'interesting'.
Ready? What I found is this:
A) There is no demonstrable connection between rs234706 and cognition.
Folate metabolism genes, vegetable intake and renal cancer risk in central Europe, by Moore et al, 2008 further reiterates the point made in the previous study.
D) There is no demonstrable link between rs234706 and angiographic venospasm.
A) Rs234706 is upregulatory for the CBS gene, not downregulatory.
B) There's a good likelihood that it's not strongly upregulatory. Why do I say so? Because if it were, you'd see significantly reduced cancer risk, and reduced risk of other illnesses where low function of CBS is a risk factor, and we don't see that here.
C) Rs234706 appears to keep cholesterol low, at least in a very particular patient group where hypercholesterolemia is a concern.
D) Final conclusion: if you have the homozygous CBS C699T 'mutation', aren't you lucky?
8) And what is the Yasko consensus regarding rs234706?
She says "it's the most severe" homozygous recessive SNP in regards to the CBS gene, and that people who have it need extra ammonia scavengers, charcoal, etc.
I'M NOT AGAINST THE PROTOCOL, GUYS, CALL OFF THE DOGS, I'M JUST INTERESTED, OKAY? THIS IS FOR SCIENCE.
You should also know in no way did I pick/choose articles to support one particular point of view. I wasn't even aware of what they were 'proving' in the article when I selected it; I solely used the selection criteria as outlined above. I admit I didn't scroll through all 130 articles - just under half of them, actually - but I figured the point was pretty darned clear. As always, you are welcome to go out and do your own research, and if you've learned anything at all by reading this, it should be that I encourage you to do so.
I also really, really don't care what you're doing, so long as it works for you. Plus, I don't disagree with the Yasko protocol, for the most part.
I should say that many of the articles I read discussed SNPs that affected CBS that were detrimental to its function, and a few articles that discussed MTHFR SNPs in conjunction that also had negative effects. I'm also not saying that no SNPs are significant! Just that it's necessary to take it all on a case-by-case basis. Which can be maddening, but is important.
And now I'm too tired, so I'm going to post this soonish.
One last thing, though:
9) But JaimeS, I can't do this for every SNP.
I know. I know.
This took me one day to write. But it was intensive. And intense. And I can't actually imagine doing it 600,000 times. Or even the 600 times it might take to grab my 'red' SNPs.
But I probably will.
There's something in me that can't simply accept what I'm told without examining it first. And that habit has saved me again and again in this illness. One does not throw away a tool in one's arsenal, even when it's cumbersome.
Still, and here's the takeaway point: you don't have to do it for all of them - just the ones that are controversial, or where you feel their significance is unclear.
And now, good night, all!
-J
[Edited 5/1/2015, primarily for spelling and grammar errors.]
<---Start the previous series
<---------Start the first series
You've gathered all the information you can. You've used some of the services to try and interpret your data. What now?
Let's face the problem head-on. As ME patients, we don't have a lot of focus. It's hard to devote any kind of energy to anything, and that includes intellectual energies. I'm a devoted author who loves to write, but it's become very hard for me to work on anything long-term because juggling that many variables eventually gets to feel like work; it loses its entertainment value and I'm left feeling like writing fantasy is a job I have to push through.
Short articles like this are usually within my grasp. As you'll see by looking at my closing statements, I usually just stop when I get to feeling like that and say, "see you next time!" We can't always do that in other arenas of our lives. I've always had a lot of difficulty stopping a job partway through and picking it up coherently, later, which is why my best stories are always the ones completed within a few days. I believe this is an aspect of my personality rather than something ME has done to my brains.
So when you're handed a list of SNPs in the hundreds of thousands - or at least, when I am - my brain looks at that long list and fizzles out like a candle in a high wind.
So it's tempting to look at the work that someone else has done and say, "well, I guess that seems reasonable." It's also very tempting to simplify and say that any negative SNP means disaster, or that none of it has significant impact at all. If it's some one way and some the other, and sometimes SNP X only has a significant impact if SNP Y is also involved, it gets complicated.
Let's take our good old friend, CBS C699T. After all, I have the dreaded red box for that SNP, so it's in my best interest to find out what kind of impact that will have on my life. I'm going to show you the process I used to determine the meaningfulness of this SNP.
1) Go to pubmed, scroll down the 'Search Entrez' box until it says 'SNP' and enter the SNP ID number of your SNP (that rs#### thingie). When you hit enter, a list of several different links will pop up with this rs#. Click the SNP ID number that says 'Homo sapiens' next to it (the first link.) This will pop up:
NCBI has an enormous amount of information regarding each SNP, but it's a little hard to wade through. We're gonna take this one step at a time, though.
Let's take a closer look at the middle column that says "allele":
2) If you click where it says 'ClinVar', it will take you here:
This is more detailed information regarding this SNP. Look on the right side:
It says that the clinical significance is benign, and lists no conditions associated with this SNP. The two stars say that more than one researcher has provided evidence that this SNP is benign. So you don't have to worry about this SNP, at least as far as NCBI is concerned.
Okay. Let's assume we don't trust their bloody medical opinions. Frankly, if you have utmost faith in The Man and the Establishment and you have this illness, you haven't been paying enough attention.
It does say that the cystathionine-beta-synthase gene is affected. Let's look into that a little more.
3) Click where it says 'Gene' next to CBS's name. This will take you to a page devoted to the CBS gene. If you scroll down this new page:
______________________________________________
Display Settings:
Send to:
CBS cystathionine-beta-synthase [ Homo sapiens (human) ]
Gene ID: 875, updated on 26-Apr-2015
Summary
Official Symbol
CBSprovided by HGNC
Official Full Name
cystathionine-beta-synthaseprovided by HGNC
Primary source
HGNC:HGNC:1550
See related
Ensembl:ENSG00000160200; HPRD:01994; MIM:613381; Vega:OTTHUMG00000086834
Gene type
protein coding
RefSeq status
REVIEWED
____________________________________________________
Display Settings:
Send to:
CBS cystathionine-beta-synthase [ Homo sapiens (human) ]
Gene ID: 875, updated on 26-Apr-2015
Summary
Official Symbol
CBSprovided by HGNC
Official Full Name
cystathionine-beta-synthaseprovided by HGNC
Primary source
HGNC:HGNC:1550
See related
Ensembl:ENSG00000160200; HPRD:01994; MIM:613381; Vega:OTTHUMG00000086834
Gene type
protein coding
RefSeq status
REVIEWED
____________________________________________________
You can find articles about the CBS gene like there's no tomorrow. 136 regarding references into functions as of this posting! That is pretty darned impressive, but not possible to read through. You can, however, open up all of those by going here:
4) Open up 'see all' at the bottom of the page. Then, you won't just see the articles, but summaries of what each of them convey. This makes them very quick to scroll through and see if you can find anything interesting.
Look at those handy little summaries on the left! How lovely of them. <3
5) Determine what looks 'interesting'.
- Does it mention your SNP by name? (Ah, I discovered yet another naming convention: CBS C699T is also called Ex9+33C>T, ugh).
- Does it discuss the SNPs that affect the gene in a way that hints it may contain info regarding your SNP?
- Is full-text available? Just because the researchers say their study demonstrated X doesn't make it true.
- Is the study a larger study, or is it student's t-test material? Twenty people is a pilot study, and may show directions for future research, but it doesn't demonstrate squat.
- In order from most to least reputable: done on humans; done on mammals; done on a bacterium with the SNP genetically inserted; not in a living thing at all.
Ready? What I found is this:
A) There is no demonstrable connection between rs234706 and cognition.
A Genetic association of cognitive ability and cognitive ageing using 325 markers for 109 genes associated with oxidative stress or cognition by Harris et al, 2007. They looked at the Moray House Test, Raven's Progressive test, Verbal Fluency, and Logical Memory. No dice whatsoever. BIG study, too.
B) There is no demonstrable connection between rs234706 and lymphoma progression.
Gene-nutrient interactions among determinants of folate and one-carbon metabolism on the risk of non-Hodgkins lymphoma: NCI-SEER Case-Control study, by Lim et al, 2007 found that patients with poor one-carbon metabolism were more likely to progress rapidly in non-Hodgkins lymphoma. They found that if you had rs1801181, yes, you were sicker quicker. However, this was not the case with rs234706, which seemed to have no effect. This implies (but does NOT unequivocally state) that this SNP does not negatively impact your B6 or methionine levels - or positively affect them.
C) There is no demonstrable link between rs234706 and renal cancer.
Folate metabolism genes, vegetable intake and renal cancer risk in central Europe, by Moore et al, 2008 further reiterates the point made in the previous study.
D) There is no demonstrable link between rs234706 and angiographic venospasm.
Gain-of-function polymorphisms of cystathione B-synthase and delayed cerebral ischemia following aneurysmal subarachnoid hemorrahges, by Brobelny et al, 2011, found no link between rs234706 and angiographic vasospasm - but they thought the SNP might help, not harm. Note that the above article, and everyone else who mentions it one way or the other, identifies rs234706 as a gain of function polymorphism. That is, it does not attenuate the CBS gene; it amplifies it.
E) Rs234706 is protective against hypercholesterolemia in superficial and deep vein thromboses.
Homocysteine levels and polymorphisms of MTHFR and CBS genes in Columbian patients with superficial and deep vein thrombosis, by Ayala et al, 2010 show that CBS C699T is protective AGAINST both superficial and deep vein thromboses.
This was actually a pretty small study, rather (only 33 people), so YMMV. Still, it's certainly promising. They excluded people with diagnoses that might have messed up their results, such as lupus, anti-phospholipid syndrome, and cancer, and found that people who carried rs234706 had significantly lower cholesterol than their counterparts.
F) Chemical chaperones such as aminolevulinic acid, betaine, and glycerol may help in actual CBS deficiency (which CBS C699T 'mutations' do NOT cause).This was actually a pretty small study, rather (only 33 people), so YMMV. Still, it's certainly promising. They excluded people with diagnoses that might have messed up their results, such as lupus, anti-phospholipid syndrome, and cancer, and found that people who carried rs234706 had significantly lower cholesterol than their counterparts.
So saith Restoring assembly and activity of cystathionine B-synthase mutants by ligands and chemical chaperones by Kopecka et al, 2011. Keep in mind that this refers to actual mutations rather than unruly SNPs, but if my reasoning is correct, this would help anyone with partial malfunction of the CBS gene. ALA (aminolevulinic acid, not alpha-lipoic) is apparently most helpful in the regulation of hemes, and the strongest, while betaine is weaker, and a methyl donor. Glycerol is the chemical chaperone referred to in the title and has more general applications, helping with a wide array of mutations and malfunctions than either of the other two. They found that SAM, L-serine, and taurine didn't help much.
Of course, this isn't for you if you have rs234706, but another of the CBS red boxes, or several. Because as should be becoming more and more clear, if you have rs234706, you don't have a loss of function.
7) And Conclusions:Of course, this isn't for you if you have rs234706, but another of the CBS red boxes, or several. Because as should be becoming more and more clear, if you have rs234706, you don't have a loss of function.
A) Rs234706 is upregulatory for the CBS gene, not downregulatory.
B) There's a good likelihood that it's not strongly upregulatory. Why do I say so? Because if it were, you'd see significantly reduced cancer risk, and reduced risk of other illnesses where low function of CBS is a risk factor, and we don't see that here.
C) Rs234706 appears to keep cholesterol low, at least in a very particular patient group where hypercholesterolemia is a concern.
D) Final conclusion: if you have the homozygous CBS C699T 'mutation', aren't you lucky?
8) And what is the Yasko consensus regarding rs234706?
She says "it's the most severe" homozygous recessive SNP in regards to the CBS gene, and that people who have it need extra ammonia scavengers, charcoal, etc.
I'M NOT AGAINST THE PROTOCOL, GUYS, CALL OFF THE DOGS, I'M JUST INTERESTED, OKAY? THIS IS FOR SCIENCE.
You should also know in no way did I pick/choose articles to support one particular point of view. I wasn't even aware of what they were 'proving' in the article when I selected it; I solely used the selection criteria as outlined above. I admit I didn't scroll through all 130 articles - just under half of them, actually - but I figured the point was pretty darned clear. As always, you are welcome to go out and do your own research, and if you've learned anything at all by reading this, it should be that I encourage you to do so.
I also really, really don't care what you're doing, so long as it works for you. Plus, I don't disagree with the Yasko protocol, for the most part.
I should say that many of the articles I read discussed SNPs that affected CBS that were detrimental to its function, and a few articles that discussed MTHFR SNPs in conjunction that also had negative effects. I'm also not saying that no SNPs are significant! Just that it's necessary to take it all on a case-by-case basis. Which can be maddening, but is important.
And now I'm too tired, so I'm going to post this soonish.
One last thing, though:
9) But JaimeS, I can't do this for every SNP.
I know. I know.
This took me one day to write. But it was intensive. And intense. And I can't actually imagine doing it 600,000 times. Or even the 600 times it might take to grab my 'red' SNPs.
But I probably will.
There's something in me that can't simply accept what I'm told without examining it first. And that habit has saved me again and again in this illness. One does not throw away a tool in one's arsenal, even when it's cumbersome.
Still, and here's the takeaway point: you don't have to do it for all of them - just the ones that are controversial, or where you feel their significance is unclear.
And now, good night, all!
-J
[Edited 5/1/2015, primarily for spelling and grammar errors.]
<---Start the previous series
<---------Start the first series