You are using an out of date browser. It may not display this or other websites correctly.
You should upgrade or use an alternative browser.
You should upgrade or use an alternative browser.
"no call" for all GSTT1 genes. What does this mean?
- Thread starter kday
- Start date
Ok, do I not have the gene? It says enzyme activity is absent in 20% of caucasians. Does this mean I wouldn't have any SNPs. Do I have a gene deletion? I'll be the first to say that I don't really understand this.
Does anybody else get "no call" for all GSTT1 SNPs?
Does anybody else get "no call" for all GSTT1 SNPs?
Glutathione S-transferase theta enzyme activity involved in the metabolism of toxic compounds is absent in approximately 20% of Caucasians due to a homozygous deletion of GSTT1 (*0/0). Because the exact manner of the GSTT1 deletion was unknown, current genotyping of GSTT1 was limited to detect the presence versus complete absence of the gene by a GSTT1-specific polymerase chain reaction (PCR). Thus, heterozygous (*A/0) and homozygous (*A/A) samples could not be discriminated. We have characterized the boundaries of the deletion of the human glutathione S-transferase theta (GSTT1) gene: PCR mapping and sequencing revealed a 54251 bp fragment including GSTT1 to be deleted from chromosome 22, most likely by a homologous recombination event between two highly homologous sequence stretches that flank GSTT1. Based on the knowledge of the GSTT1*0 region, a PCR assay was devised for unambiguous discrimination of homozygously deleted (*0/0), heterozygously (*A/0) and homozygously GSTT1 carrying (*A/A) individuals. Genotyping of 180 samples of a Caucasian population revealed that the deletion consists of one defined allele, whose distribution in the population fits the Hardy-Weinberg equilibrium with observed 20% *0/0, 46% *A/0 and 34% *A/A individuals. The number of GSTT1*A alleles detected by this procedure correlated highly significant with the enzyme activity in erythrocytes. Genotype-phenotype comparisons demonstrated a codominant type of inheritance by a gene-dose effect: samples with two active alleles expressed a statistically significant higher enzymatic activity compared to those with one null allele (P < 0.0001, ANOVA).
GSTM1 deletion and/or GSTP1 polymorphisms seem to be more common among ME/CFS people according to Genovations (DetoxiGenomic) gene tests that I have received from people. Maybe it is the same with the GSTT1 . It would not surprise me.
Are the GSTM1 and GSTP1 genes also analysed in the test from 23andme
?
Are the GSTM1 and GSTP1 genes also analysed in the test from 23andme
?
Yes, however, all of the SNPs are there for all 40 of GSTM1 and GSTP1 SNPs. 23andMe does not report my GSTT1 as a deletion (they say they don't regularly keep track of deletions), but instead says "no call" for all SNPs. However, I think it's logical that it is probably a deletion. But I sent in a support ticket asking them if it could be.
Yes, however, all of the SNPs are there for all 40 of GSTM1 and GSTP1 SNPs. 23andMe does not report my GSTT1 as a deletion (they say they don't regularly keep track of deletions), but instead says "no call" for all SNPs. However, I think it's logical that it is probably a deletion. But I sent in a support ticket asking them if it could be.
Hi Kday,
Did you get any reply from 23andme?
- Messages
- 60
- Location
- Michigan
Yes, I've been waiting to see if 23andme got back with you on this issue. I didn't get a no-call for this, but just a straight up deletion I think.
Sea
Senior Member
- Messages
- 1,286
- Location
- NSW Australia
I have a "no call" for 12 out of the 14 snps tested for GSTT1. I've just assumed it's a deletion since it's not uncommon for there to be significant deletions in this gene.
- Messages
- 55
- Location
- Auckland, NZ
Same here:
GSTT1 - Absent*
*It is likely that you are missing this gene and have a deletion. However, since this type of testing isn't technically designed to detect deletions, it isn't with 100% certainty.
GSTT1 - Absent*
*It is likely that you are missing this gene and have a deletion. However, since this type of testing isn't technically designed to detect deletions, it isn't with 100% certainty.
Critterina
Senior Member
- Messages
- 1,238
- Location
- Arizona, USA
Me, too. I have rs11550605 TT and rs11550605 is AA. The rest are no call. Sea, Are these the same ones you have?I have a "no call" for 12 out of the 14 snps tested for GSTT1. I've just assumed it's a deletion since it's not uncommon for there to be significant deletions in this gene.
In areas where I have an insertion or deletion, I get an I or D in my report, instead of A,C,G, or T.
Sea
Senior Member
- Messages
- 1,286
- Location
- NSW Australia
Me, too. I have rs11550605 TT and rs11550605 is AA. The rest are no call. Sea, Are these the same ones you have?
In areas where I have an insertion or deletion, I get an I or D in my report, instead of A,C,G, or T.
Yes, I have the same two snps - assuming you mean 11550605 TT and 11550606 AA. According to SNPedia these are both the ancestral allele, but according to the human gene compendium the 11550606 AA is the variant and is a missense mutation. I don't know which one is more reliable.
http://www.genecards.org/cgi-bin/carddisp.pl?gene=GSTT1&snp=133#snp
I got my 23andme results in March this year. How long have you had your results? They may have changed the way they report. I remember reading somewhere on their site that they would go back to no call results and update them as they got new information.
I have some II and DD elsewhere in my report
Critterina
Senior Member
- Messages
- 1,238
- Location
- Arizona, USA
I got my results late July (~26th). I just poked into 'browse' to respond to this post, though, so my no-calls were from their website the day I made the post.How long have you had your results?
aquariusgirl
Senior Member
- Messages
- 1,736
no call here too, except for the 11550606 AA. Rich would be all over this...god bless him...
- Messages
- 19
- Location
- Illinois USA
I got tested in July 2014 with the V4 chip. I have no call on all except 3 of the SNPs 23andMe shows for GSTT1.
rs1130990: CC (less common than G but not rare)
rs11550605: GT (occurs <.05%)
rs2266636: CT (appears to occur < 1% looking at openSNP, but didn't come up in rare allele program, unsure)
Here is a resource I hadn't seen before: Wolfram
rs1130990: CC (less common than G but not rare)
rs11550605: GT (occurs <.05%)
rs2266636: CT (appears to occur < 1% looking at openSNP, but didn't come up in rare allele program, unsure)
Here is a resource I hadn't seen before: Wolfram
Last edited:
If you have any SNPs on GSTT1, then you have the gene.I got tested in July 2014 with the V4 chip. I have no call on all except 3 of the SNPs 23andMe shows for GSTT1.
23andMe lists a lot of SNPs on that gene, but usually just tests a few for most customers, hence they show a lot of no call results.
This is weird. Why would this happen? It kinda freaked me out a bit.
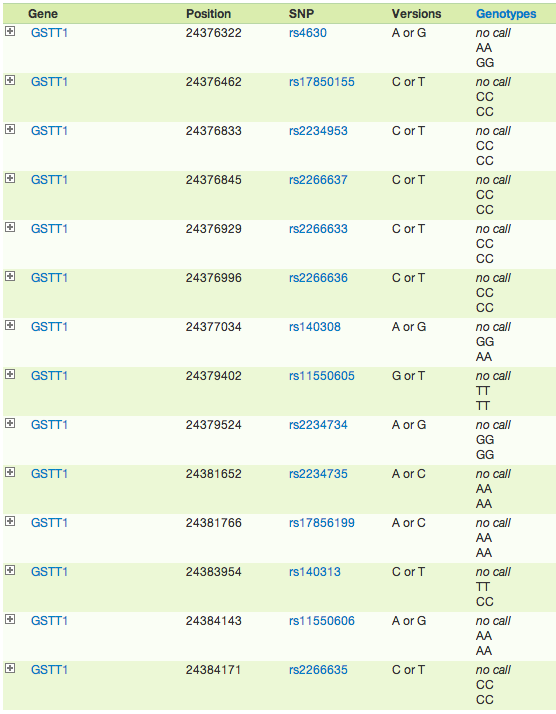
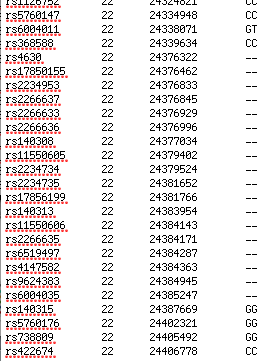
What report was the original poster using that showed "No Call" for different genes? The 23andme raw data browser does not show that.
