So, ladies and gents, I haven't received any results back from de Meirlier's offices yet, but two topics have been a-brewing in my brain meanwhile. I figured I'd give the tougher one of those two a shot.
The 'tougher' issue is SNPs. I'm going to write this up as a primer, so that anyone who comes in as clueless as I was can have a place to start.
What is a SNP?
SNP stands for 'single nucleotide polymorphism'. Let's break that down: single, meaning one; nucleotide, meaning a particular chemical, which I will discuss below; and polymorphism, where poly means many and morph means shape, so many-shaped.
Nucleotides are parts of DNA. All nucleotides consist of a sugar (ribose or deoxyribose in our case), a base (adenine, guanine, cytosine or thymine in our case), and at least one phosphate group.
So, strung together, 'SNP' refers to a chemical part of our DNA that can vary between members of the same species, i.e. you vs me. What makes me look different from you? What makes my gut motility faster or slower? Why do I need a high-dose Vitamin B12 pill, but the same supplement gives you symptoms of toxicity? These variations have to do with which bases are present in the nucleotide. My DNA might have two adenines paired together, but you might have an adenine and a cytosine. This is why, while neither of us have a mutation, we still appear different, respond differently to same environment, and respond differently to the same chemicals.
Let's say that on the SNP called "CBS C699T" I have two thymines. That makes my alleles 'TT'. You might have something different, like 'GG'. Let's talk about what that means in practical terms.
In order to do that, we have to discuss bases in greater detail, so get ready to flash back to high school biology! <<<TIIIIMMME WAARRRP>>>>
High School Biology!
Okay. Whew! We're here.
So there are four bases, as previously stated:
Adenine (abbreviated 'A')
Thymine (abbreviated 'T')
Cytosine (C)
Guanine (G)
Each of these bases pair up with a partner, one on either side of the double-helix of the DNA strand.
Adenine pairs with thymine
Guanine pairs with cytosine
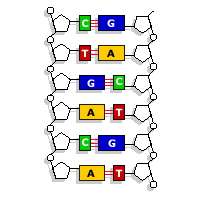
(From BBC News, 2006)
For this reason, you might see my 'TT' for CBS C699T listed as 'AA', depending on whose report we're looking at. In this case, the other side of the DNA strand was analyzed. Although adenine and thymine are NOT identical, if one is on one half of the DNA double-strand, its 'partner' must be on the other side. So if an analysis program tells you you have AA for a SNP, then the other half of the strand must be TT. (Thanks to @Valentijn for pointing that out!)
In the general population, for the SNP CBS C699T, there are three possible combinations:
CC (which could also be written as 'GG')
TT (which could also be written as 'AA')
CT (which could also be written as 'GA')
It's important to note that when you have two of the same bases in a row (CC or TT), we call this homozygous ('homo' means 'same'), and where we've got a mix-n-match (CT, for example), we call this heterozygous ('hetero' means 'different'.)
When a geneticist names a SNP "C699T" they are giving you the commoner base first, followed by the rarer base. That means that often, CC is the most common, followed by CT, and finally TT would be the rarest or most unusual. [Edit: although, as @Valentijn has pointed, out, the heterozygous is sometimes the most common, if the 'rarer' allele's frequency approaches 50%. Thanks, V!] So far as I am aware, all SNP names follow this simple pattern, making it easy to tell the rarer and more common combos apart, just by looking. The 'CBS' refers to what the SNP (partially) regulates: in this case, cystathionine beta synthase, which plays a role in the methylation (one-carbon) cycle.
Even if you have TT however, it is not, technically speaking a mutation. A mutation occurs when there is a 'mistake' in the transcription or translation of DNA: that is, you display a trait that neither of your parents possessed or carried, because in the process of copying your DNA in the womb, Mistakes Were Made. That is likely not what has happened, here. Instead, one or both of your parents have passed the rarer base(s) down to you, in the totally normal manner.
This stricter definition of the word is not held by everyone. Sometimes it is viewed as a difference in DNA that is not carried or expressed in most organisms of the same species, and you can find both definitions if you look online.
However, I stick to the definition I learned in my many biology classes over the years.
Getting Your SNPs Analyzed
23 and me:
The cheapest way is by going through 23andme. Although the site only analyzes approximately 10% of disease-causing SNPs, it also only costs $100, and you get neat little graphics and ancestry information (I never knew my Dad's side had East Asian ancestors!) The downside is that analysis on your own can be confusing and frustrating.
You will receive a chunk of raw data that contains about 600,000 SNPs. You can use Valentjin's program to automatically yank out the SNPs that are rarest (and therefore have the highest likelihood of being an issue). Her program will yank out SNPs that only 1% or fewer individuals carry, and create a separate file for those that 10% or fewer individuals carry. That's when it starts to get interesting.
When I yanked out my 1%ers, there were over 100 SNPs. (This may or may not have something to do with the fact that I am apparently 48% Ashkenazi Jewish. In case you haven't heard, we have higher incidences of several genetic illnesses that are extremely uncommon in the general population.)
I had a mistake here regarding how SNPs with a 1% prevalence should comprise 1% of 600,000 SNPs (i.e., 6,000 SNPs). Here's why that's incorrect:
Most SNPs don't have any allele pairs with a 1% prevalence. Generally, all three possibilities are rather likely. Take CBS C699T, for example (since we have been, all along!) In my population, the prevalence is approximately:
CC: 60%
CT: 30%
TT: 10%
So the likelihood that I'll have a 1% allele for that? It's zero, because 1% or less is not an option in the case of that SNP.
This rules out an enormous number of SNPs - the bulk of them. For the remaining SNPs, where it is POSSIBLE that there is a 1% prevalence, you have a 1% chance of having each one.
So the issue is one of confused variables: how likely the SNP is to have a 1% or lower prevalence allele pair vs how likely you are to have one of the SNPs that fit that criteria.
If you have insurance coverage, Illumina will run your entire exome, which will cost significantly more but will provide a far greater amount of information.
There are several other programs that you might choose to run your file through. None of the following cost more than $40.
Promethease:
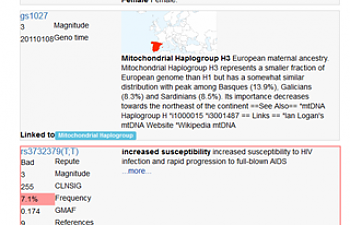
Promethease is quite possibly the best of these. It gives you a real analysis of your SNPs and some more general information as well:
Note that it tells me where my mother's ancestors came from (my haplogroup, H3), that I'm a lady (er, thanks), and then it gets to the SNPs it thinks are most important. Apparently, I really shouldn't contract HIV (good to know).
Promethease also tells you whether the SNP is considered a good thing, a bad thing, or nothing (for example, it cheerfully informed me that I am 70% more likely to think that cilantro tastes like soap. IT DOES.) It indicates this with a red border, a green border, or a neutral-colored border, but it also says BAD or GOOD at the top left, just in case we missed it.
It lists the SNP ID (a number-letter string that typically begins with 'rs'), discusses its significance in the larger space on the right, and, in the black box below, discusses in more detail other conditions this SNP may be related to. Best of all, it links out, so you can easily find the scholarly article that first discussed the significance of the SNP and actually read it. This is brilliant.
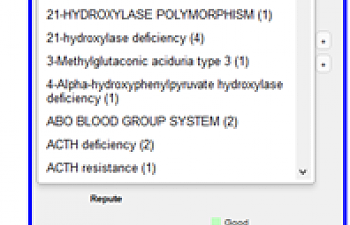
Quite possibly best of all, if there is something in particular that is concerning to you, you can use the drop-down box off to the side to search for SNPs related to particular topics/ illness states:
Moreover, you can see a little pie chart at the bottom that rates how reputable they find the connection between the SNP and the condition. More on this sort of thing later.
There is a LOT more that Promethease can do, but that's it for now - you'll have to explore on your own if you decide it's for you.
SNPedia:
SNPedia is a great resource for those who are more research-minded. Find the SNP on your 'rare' list (or any SNP that interests you) and plug its SNP ID ( rs###### ) into the search box. Hit enter and wait.
And wait.
For some reason, it'll take awhile.
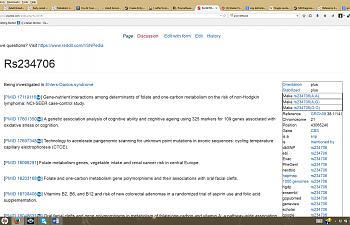
Then, something like this will pop up:
For purposes of consistency, I'm continuing to use CBS C699T as my 'demo SNP'. At the top of the page is its SNP ID, rs234706. Right below that, it says, "being investigated in Ehlers-Danlos syndrome". This doesn't mean anything, just that someone is doing a study. Below that is the good stuff.
Each link starts with 'PMD' because it links directly to an article on pubmed that specifically mentions this SNP. Next to that link, some helpful soul has listed the name of the article in question, and sometimes a quick blurb regarding the article's conclusions.
You'll find that, in general, more certain conclusions are discussed as such. In such cases, the connection will be listed right below the SNP ID number with some degree of finality. This, however, is rather rare.
MTHFR Support:
Some of you may have heard of Amy Yasko and her methylation protocol. Yasko was originally focused on treating autism, but later expanded her treatment protocol to other illnesses such as chronic fatigue syndrome and ME. Her main focus were errors in one-carbon metabolism, also known as the methylation cycle. When you hear people on PR discussing 'ammonia toxicity', homocysteine, and depleted B Vitamins, especially in connection with heavy metal toxicity and liver clearance, you are hearing about Yasko's protocol and ideas.
YMMV, but my stance on Yasko is a bit wobbly. Some of her science does not appear to be substantiated, but I feel like generally her protocol is on the right track.
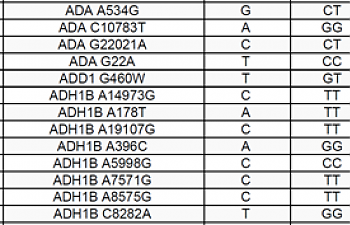
Regardless, if you really want an analysis of the SNPs that directly relate to metabolism in general, and the methylation cycle in particular, you want Sterling's app on the MTHFR Support page, which will analyze your SNPs and spit out something like this:
This will go on for pages, but it will be separated into categories like 'IgE', 'Alzheimer's/Cardio/Lipid', and the biggie, 'Methylation & Methionine/Homocysteine Pathways'. The green represents where you have the commonest base pair. The yellow represents where you are heterozygous for the more and less common forms, and red is when you have the least common/pathogenic allele pair.
pubmed
Finally, there's pubmed.gov, that lovely repository of marvelous scientific articles from all across the globe. It's most efficacious to search for the SNP ID # or, if you want to look more generally, you can use the gene itself, spelled out (cystathionine beta synthase).
...but What Does It All Mean?
Guys, here's the tough part. Unless you get one of those 'definite' statements at the top of SNPedia, it's really tough to tell how affected you are by your less usual SNPs.
Keep this in mind especially in regard to the Yasko-associated information. Sometimes, one defect is enough to significantly affect metabolism, but not usually. The thing about the human body is that it has backups on backups. There are six SNPs that I was tested for which are associated with CBS. I only have one 'red' box for CBS. I probably shouldn't panic about CBS, in this case!
One of the most useful things that SNPedia shows you is prevalence of your SNP. If 25% of the population shares your 'rarer' SNP, it's also likely not causing you any major difficulties. Promethease, too, shows prevalence as well as how sure they feel that the SNP has negative consequences.
To Be Continued....
<--Start the previous series
<-------Start the first series
The 'tougher' issue is SNPs. I'm going to write this up as a primer, so that anyone who comes in as clueless as I was can have a place to start.
What is a SNP?
SNP stands for 'single nucleotide polymorphism'. Let's break that down: single, meaning one; nucleotide, meaning a particular chemical, which I will discuss below; and polymorphism, where poly means many and morph means shape, so many-shaped.
Nucleotides are parts of DNA. All nucleotides consist of a sugar (ribose or deoxyribose in our case), a base (adenine, guanine, cytosine or thymine in our case), and at least one phosphate group.
So, strung together, 'SNP' refers to a chemical part of our DNA that can vary between members of the same species, i.e. you vs me. What makes me look different from you? What makes my gut motility faster or slower? Why do I need a high-dose Vitamin B12 pill, but the same supplement gives you symptoms of toxicity? These variations have to do with which bases are present in the nucleotide. My DNA might have two adenines paired together, but you might have an adenine and a cytosine. This is why, while neither of us have a mutation, we still appear different, respond differently to same environment, and respond differently to the same chemicals.
Let's say that on the SNP called "CBS C699T" I have two thymines. That makes my alleles 'TT'. You might have something different, like 'GG'. Let's talk about what that means in practical terms.
In order to do that, we have to discuss bases in greater detail, so get ready to flash back to high school biology! <<<TIIIIMMME WAARRRP>>>>
High School Biology!
Okay. Whew! We're here.
So there are four bases, as previously stated:
Adenine (abbreviated 'A')
Thymine (abbreviated 'T')
Cytosine (C)
Guanine (G)
Each of these bases pair up with a partner, one on either side of the double-helix of the DNA strand.
Adenine pairs with thymine
Guanine pairs with cytosine

(From BBC News, 2006)
For this reason, you might see my 'TT' for CBS C699T listed as 'AA', depending on whose report we're looking at. In this case, the other side of the DNA strand was analyzed. Although adenine and thymine are NOT identical, if one is on one half of the DNA double-strand, its 'partner' must be on the other side. So if an analysis program tells you you have AA for a SNP, then the other half of the strand must be TT. (Thanks to @Valentijn for pointing that out!)
In the general population, for the SNP CBS C699T, there are three possible combinations:
CC (which could also be written as 'GG')
TT (which could also be written as 'AA')
CT (which could also be written as 'GA')
It's important to note that when you have two of the same bases in a row (CC or TT), we call this homozygous ('homo' means 'same'), and where we've got a mix-n-match (CT, for example), we call this heterozygous ('hetero' means 'different'.)
When a geneticist names a SNP "C699T" they are giving you the commoner base first, followed by the rarer base. That means that often, CC is the most common, followed by CT, and finally TT would be the rarest or most unusual. [Edit: although, as @Valentijn has pointed, out, the heterozygous is sometimes the most common, if the 'rarer' allele's frequency approaches 50%. Thanks, V!] So far as I am aware, all SNP names follow this simple pattern, making it easy to tell the rarer and more common combos apart, just by looking. The 'CBS' refers to what the SNP (partially) regulates: in this case, cystathionine beta synthase, which plays a role in the methylation (one-carbon) cycle.
Even if you have TT however, it is not, technically speaking a mutation. A mutation occurs when there is a 'mistake' in the transcription or translation of DNA: that is, you display a trait that neither of your parents possessed or carried, because in the process of copying your DNA in the womb, Mistakes Were Made. That is likely not what has happened, here. Instead, one or both of your parents have passed the rarer base(s) down to you, in the totally normal manner.
This stricter definition of the word is not held by everyone. Sometimes it is viewed as a difference in DNA that is not carried or expressed in most organisms of the same species, and you can find both definitions if you look online.
However, I stick to the definition I learned in my many biology classes over the years.
Getting Your SNPs Analyzed
23 and me:
The cheapest way is by going through 23andme. Although the site only analyzes approximately 10% of disease-causing SNPs, it also only costs $100, and you get neat little graphics and ancestry information (I never knew my Dad's side had East Asian ancestors!) The downside is that analysis on your own can be confusing and frustrating.
You will receive a chunk of raw data that contains about 600,000 SNPs. You can use Valentjin's program to automatically yank out the SNPs that are rarest (and therefore have the highest likelihood of being an issue). Her program will yank out SNPs that only 1% or fewer individuals carry, and create a separate file for those that 10% or fewer individuals carry. That's when it starts to get interesting.
When I yanked out my 1%ers, there were over 100 SNPs. (This may or may not have something to do with the fact that I am apparently 48% Ashkenazi Jewish. In case you haven't heard, we have higher incidences of several genetic illnesses that are extremely uncommon in the general population.)
I had a mistake here regarding how SNPs with a 1% prevalence should comprise 1% of 600,000 SNPs (i.e., 6,000 SNPs). Here's why that's incorrect:
Most SNPs don't have any allele pairs with a 1% prevalence. Generally, all three possibilities are rather likely. Take CBS C699T, for example (since we have been, all along!) In my population, the prevalence is approximately:
CC: 60%
CT: 30%
TT: 10%
So the likelihood that I'll have a 1% allele for that? It's zero, because 1% or less is not an option in the case of that SNP.
This rules out an enormous number of SNPs - the bulk of them. For the remaining SNPs, where it is POSSIBLE that there is a 1% prevalence, you have a 1% chance of having each one.
So the issue is one of confused variables: how likely the SNP is to have a 1% or lower prevalence allele pair vs how likely you are to have one of the SNPs that fit that criteria.
If you have insurance coverage, Illumina will run your entire exome, which will cost significantly more but will provide a far greater amount of information.
There are several other programs that you might choose to run your file through. None of the following cost more than $40.
Promethease:
Promethease is quite possibly the best of these. It gives you a real analysis of your SNPs and some more general information as well:
Note that it tells me where my mother's ancestors came from (my haplogroup, H3), that I'm a lady (er, thanks), and then it gets to the SNPs it thinks are most important. Apparently, I really shouldn't contract HIV (good to know).
Promethease also tells you whether the SNP is considered a good thing, a bad thing, or nothing (for example, it cheerfully informed me that I am 70% more likely to think that cilantro tastes like soap. IT DOES.) It indicates this with a red border, a green border, or a neutral-colored border, but it also says BAD or GOOD at the top left, just in case we missed it.
It lists the SNP ID (a number-letter string that typically begins with 'rs'), discusses its significance in the larger space on the right, and, in the black box below, discusses in more detail other conditions this SNP may be related to. Best of all, it links out, so you can easily find the scholarly article that first discussed the significance of the SNP and actually read it. This is brilliant.
Quite possibly best of all, if there is something in particular that is concerning to you, you can use the drop-down box off to the side to search for SNPs related to particular topics/ illness states:
Moreover, you can see a little pie chart at the bottom that rates how reputable they find the connection between the SNP and the condition. More on this sort of thing later.
There is a LOT more that Promethease can do, but that's it for now - you'll have to explore on your own if you decide it's for you.
SNPedia:
SNPedia is a great resource for those who are more research-minded. Find the SNP on your 'rare' list (or any SNP that interests you) and plug its SNP ID ( rs###### ) into the search box. Hit enter and wait.
And wait.
For some reason, it'll take awhile.
Then, something like this will pop up:
For purposes of consistency, I'm continuing to use CBS C699T as my 'demo SNP'. At the top of the page is its SNP ID, rs234706. Right below that, it says, "being investigated in Ehlers-Danlos syndrome". This doesn't mean anything, just that someone is doing a study. Below that is the good stuff.
Each link starts with 'PMD' because it links directly to an article on pubmed that specifically mentions this SNP. Next to that link, some helpful soul has listed the name of the article in question, and sometimes a quick blurb regarding the article's conclusions.
You'll find that, in general, more certain conclusions are discussed as such. In such cases, the connection will be listed right below the SNP ID number with some degree of finality. This, however, is rather rare.
MTHFR Support:
Some of you may have heard of Amy Yasko and her methylation protocol. Yasko was originally focused on treating autism, but later expanded her treatment protocol to other illnesses such as chronic fatigue syndrome and ME. Her main focus were errors in one-carbon metabolism, also known as the methylation cycle. When you hear people on PR discussing 'ammonia toxicity', homocysteine, and depleted B Vitamins, especially in connection with heavy metal toxicity and liver clearance, you are hearing about Yasko's protocol and ideas.
YMMV, but my stance on Yasko is a bit wobbly. Some of her science does not appear to be substantiated, but I feel like generally her protocol is on the right track.
Regardless, if you really want an analysis of the SNPs that directly relate to metabolism in general, and the methylation cycle in particular, you want Sterling's app on the MTHFR Support page, which will analyze your SNPs and spit out something like this:
This will go on for pages, but it will be separated into categories like 'IgE', 'Alzheimer's/Cardio/Lipid', and the biggie, 'Methylation & Methionine/Homocysteine Pathways'. The green represents where you have the commonest base pair. The yellow represents where you are heterozygous for the more and less common forms, and red is when you have the least common/pathogenic allele pair.
pubmed
Finally, there's pubmed.gov, that lovely repository of marvelous scientific articles from all across the globe. It's most efficacious to search for the SNP ID # or, if you want to look more generally, you can use the gene itself, spelled out (cystathionine beta synthase).
...but What Does It All Mean?
Guys, here's the tough part. Unless you get one of those 'definite' statements at the top of SNPedia, it's really tough to tell how affected you are by your less usual SNPs.
Keep this in mind especially in regard to the Yasko-associated information. Sometimes, one defect is enough to significantly affect metabolism, but not usually. The thing about the human body is that it has backups on backups. There are six SNPs that I was tested for which are associated with CBS. I only have one 'red' box for CBS. I probably shouldn't panic about CBS, in this case!
One of the most useful things that SNPedia shows you is prevalence of your SNP. If 25% of the population shares your 'rarer' SNP, it's also likely not causing you any major difficulties. Promethease, too, shows prevalence as well as how sure they feel that the SNP has negative consequences.
To Be Continued....
<--Start the previous series
<-------Start the first series