If you're just joining us, you may want to go back to the previous post (link at the end of the entry) because we're continuing our discussion right from where we left off:

Woo hoo! You'll see that rs61750615 "appears to be an incomplete dominant". Lo and behold, there is a citation!
If we click on it, it takes us to an article entitled, Phased Whole-Genome Genetic Risk in a Family Quartet Using a Major Allele Reference Sequence, by Dewey et al, from September 2011 in PloS Genetics.
But when I hit 'search' and type in the rs# I get nothing.
It took me awhile to figure out that the diagrams in the article do not actually get searched. Technically speaking, they're not actually on the same page as the rest of the article; there's a link to get to them. So if we click on each of the diagrams, eventually we get the following:
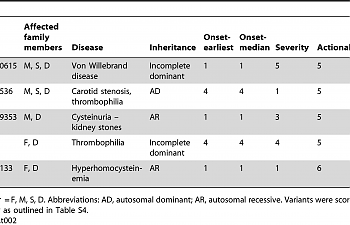
Oooh, check out the mention of the MTHFR rs1801133, which can contribute to hyperhomocysteinemia, and may add to risk of thromboembolism. That SNP should be on your methylation pages if you got them from MTHFR Support!
Apparently, in the family discussed in the article, the mother, son, and daughter had Von Willebrand disease, and rs61750615 was the only rare SNP they could find on the VWF gene. Using this evidence as well as an algorithm used to predict the hazardous nature of SNPs (the article's main thrust) the researchers conclude that this SNP is what has caused the illness. However, they believe that it is an 'incomplete dominant' trait.
For those of us who remember the TARDIS sound from the first entry, just go along with me and assume we're back in high school biology again. In classical genetics, you have a dominant trait that can 'shout over' the recessive trait. For example, I carry the allele for blue eyes from my mom, but I have my dad's brown eyes, because brown is dominant over blue for eye color. Incomplete dominance is what five-year-olds believe genetics is all about: if trait A and trait B mix, you get a trait somewhere in between. This is like breeding a red rose to a white rose to get a pink one. Or a white-and-red-stripey one. Basically, both traits get to have their say.

The example of incomplete dominance every. Single. Time.
(Wikipedia, accessed 5/4/2015)
In this case, it could mean that the patients could sometimes produce Von Willebrand factor, and other times seemed less capable of doing so. It could mean that they could produce Von Willebrand factor, but at very low levels. It could mean that they had enough for all reasonable purposes, but if there was a demand for clotting factor (i.e. a major injury), they could start failing to clot. Quite unfortunately, the researchers keep the details to themselves. The article is a genetics article, and does not discuss patient symptoms in detail.
Have you ever noted this? I sure have. When I'm looking after my own health, what's missing from the study is always the perspective of the patient.
Well, here's one, guys: the only time I ever had bleeding issues to speak of was when I had a herniated disc one summer. I also noticed that suddenly little cuts wouldn't heal, and when that time of the month rolled around... let's just say you could cross out 'time of the', and it would be more accurate.
I went to my doc, who sent me to my OBGYN, who took hormone levels and did a physical exam and came back relieved. She said that all my reproductive hormones were totally mid-range normal, and my physical didn't turn anything up either. I'd finally stopped it by pushing Black Cohosh like there was no tomorrow (a dose every 2 hours until the bleeding stopped). But we all know cellular repair was still going on a month later (or you do if you've ever had a herniated disc.) This continued for three months, after which things seemed to normalize. By then my iron and B vitamins were pretty shot, though. After asking me if I'd started passing out or 'losing time' (WHAT.) doc shrugged his shoulders as if it were just 'one of those things'. Like, oh, well! I guess we'll never really know! (Yep - it's the same fool from earlier.)
I've also noticed that my clotting rate is pretty variable, but never much to my detriment, otherwise. If I cut my finger, I've noticed sometimes it'll clot immediately, and other times I'll look at it an hour later and it'll just be closing.
So, let's say this is the only information you can get your hands on. (In my case, it is.) What else is there to do?
If you have a strong suspicion that you may have a particular condition due to a certain SNP, remember that the SNPs shouldn't be taken as the final word on anything, but used as a direction for future study and analysis. Getting a blood, urine, or stool sample analyzed for the disorder in question is the next step, if that's a possibility.
For example, Von Willebrand's Disease is diagnosable via blood tests. Unfortunately, even people who do have it (non-partially-dominantly) can sometimes come up with numbers in the normal range if they haven't bled a lot lately. Even the more official looking sites suggest you do it again three months later if it comes back negative and there's evidence of the disorder. I'm deciding whether it's worth the time, money, and energy to hunt it down like a wildebeest of the veldt.
[Edit, 3/28/16: This bleeding issue may very well be due to 'anemia of chronic infection'. After noticing that my clotting and bleeding issues seem worse when my overall illness seems fiercest, I began looking into this theory. The body produces anticoagulants in infection, which makes sense in terms of trying to clear out a wound. Discussion and citations in this thread.]
-J
Phased Whole-Genome Genetic Risk in a Family Quartet Using a Major Allele Reference Sequence
Dewey FE, Chen R, Cordero SP, Ormond KE, Caleshu C, et al. (2011) Phased Whole-Genome Genetic Risk in a Family Quartet Using a Major Allele Reference Sequence. PLoS Genet 7(9): e1002280. doi: 10.1371/journal.pgen.1002280
<----Previous in the series
<----Start the series
Woo hoo! You'll see that rs61750615 "appears to be an incomplete dominant". Lo and behold, there is a citation!
If we click on it, it takes us to an article entitled, Phased Whole-Genome Genetic Risk in a Family Quartet Using a Major Allele Reference Sequence, by Dewey et al, from September 2011 in PloS Genetics.
But when I hit 'search' and type in the rs# I get nothing.
It took me awhile to figure out that the diagrams in the article do not actually get searched. Technically speaking, they're not actually on the same page as the rest of the article; there's a link to get to them. So if we click on each of the diagrams, eventually we get the following:
Oooh, check out the mention of the MTHFR rs1801133, which can contribute to hyperhomocysteinemia, and may add to risk of thromboembolism. That SNP should be on your methylation pages if you got them from MTHFR Support!
Apparently, in the family discussed in the article, the mother, son, and daughter had Von Willebrand disease, and rs61750615 was the only rare SNP they could find on the VWF gene. Using this evidence as well as an algorithm used to predict the hazardous nature of SNPs (the article's main thrust) the researchers conclude that this SNP is what has caused the illness. However, they believe that it is an 'incomplete dominant' trait.
For those of us who remember the TARDIS sound from the first entry, just go along with me and assume we're back in high school biology again. In classical genetics, you have a dominant trait that can 'shout over' the recessive trait. For example, I carry the allele for blue eyes from my mom, but I have my dad's brown eyes, because brown is dominant over blue for eye color. Incomplete dominance is what five-year-olds believe genetics is all about: if trait A and trait B mix, you get a trait somewhere in between. This is like breeding a red rose to a white rose to get a pink one. Or a white-and-red-stripey one. Basically, both traits get to have their say.

The example of incomplete dominance every. Single. Time.
(Wikipedia, accessed 5/4/2015)
In this case, it could mean that the patients could sometimes produce Von Willebrand factor, and other times seemed less capable of doing so. It could mean that they could produce Von Willebrand factor, but at very low levels. It could mean that they had enough for all reasonable purposes, but if there was a demand for clotting factor (i.e. a major injury), they could start failing to clot. Quite unfortunately, the researchers keep the details to themselves. The article is a genetics article, and does not discuss patient symptoms in detail.
Have you ever noted this? I sure have. When I'm looking after my own health, what's missing from the study is always the perspective of the patient.
Well, here's one, guys: the only time I ever had bleeding issues to speak of was when I had a herniated disc one summer. I also noticed that suddenly little cuts wouldn't heal, and when that time of the month rolled around... let's just say you could cross out 'time of the', and it would be more accurate.
I went to my doc, who sent me to my OBGYN, who took hormone levels and did a physical exam and came back relieved. She said that all my reproductive hormones were totally mid-range normal, and my physical didn't turn anything up either. I'd finally stopped it by pushing Black Cohosh like there was no tomorrow (a dose every 2 hours until the bleeding stopped). But we all know cellular repair was still going on a month later (or you do if you've ever had a herniated disc.) This continued for three months, after which things seemed to normalize. By then my iron and B vitamins were pretty shot, though. After asking me if I'd started passing out or 'losing time' (WHAT.) doc shrugged his shoulders as if it were just 'one of those things'. Like, oh, well! I guess we'll never really know! (Yep - it's the same fool from earlier.)
I've also noticed that my clotting rate is pretty variable, but never much to my detriment, otherwise. If I cut my finger, I've noticed sometimes it'll clot immediately, and other times I'll look at it an hour later and it'll just be closing.
So, let's say this is the only information you can get your hands on. (In my case, it is.) What else is there to do?
If you have a strong suspicion that you may have a particular condition due to a certain SNP, remember that the SNPs shouldn't be taken as the final word on anything, but used as a direction for future study and analysis. Getting a blood, urine, or stool sample analyzed for the disorder in question is the next step, if that's a possibility.
For example, Von Willebrand's Disease is diagnosable via blood tests. Unfortunately, even people who do have it (non-partially-dominantly) can sometimes come up with numbers in the normal range if they haven't bled a lot lately. Even the more official looking sites suggest you do it again three months later if it comes back negative and there's evidence of the disorder. I'm deciding whether it's worth the time, money, and energy to hunt it down like a wildebeest of the veldt.
[Edit, 3/28/16: This bleeding issue may very well be due to 'anemia of chronic infection'. After noticing that my clotting and bleeding issues seem worse when my overall illness seems fiercest, I began looking into this theory. The body produces anticoagulants in infection, which makes sense in terms of trying to clear out a wound. Discussion and citations in this thread.]
-J
Phased Whole-Genome Genetic Risk in a Family Quartet Using a Major Allele Reference Sequence
Dewey FE, Chen R, Cordero SP, Ormond KE, Caleshu C, et al. (2011) Phased Whole-Genome Genetic Risk in a Family Quartet Using a Major Allele Reference Sequence. PLoS Genet 7(9): e1002280. doi: 10.1371/journal.pgen.1002280
<----Previous in the series
<----Start the series
Next in the series ---->