So, @garyfritz had a question regarding a SNP that didn't have so much information on NCBI, and it inspired me to start this entry.
There are very few SNPs that are the subject of so many studies as rs234706 (even through the worst brain fog, I will always remember that SNP!) So let's say you don't get a lot from the method that I showed you. What do you do next?
So let's say you don't get a lot from the method that I showed you. What do you do next?
Let's take Rs61750615 as our example, this time. It's one of my rarest SNPs, occurring in less that one half of 1% of the population overall.
(Sounds super-duper rare, guys, but it's still one out of about 200 people, implying that there were maybe five at my old high school with this SNP.)
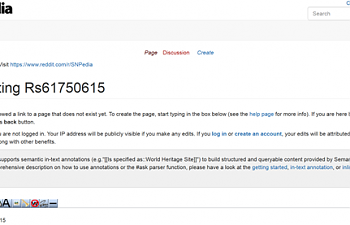
If you try to go to snpedia, this is what you'll see:
A page inviting you to please create a page for this SNP! They appear to have no information whatsoever.
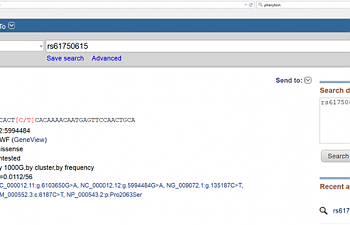
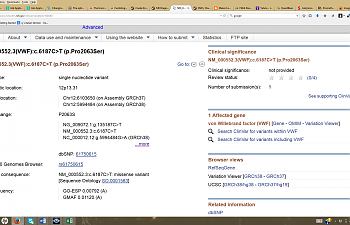
Okay, let's try our tried-and-true method described in the previous post. Go to NCBI's page, drop-down menu, select SNP, and in the search box, type the rs#. This shows up:
Well, whaddaya know? We got a little something out of it this time.
If we look where it says 'functional consequence' the page says 'missense'. This is a genetics term to avoid saying 'nonsense', but it has the same connotation.
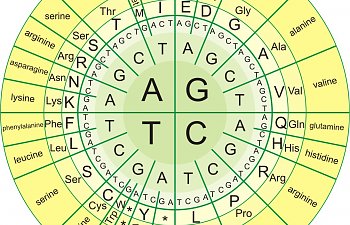
Each three bases in a row - like A, G, G, for example - is called a codon. And codons are called that because they each code for amino acids:
So, if you have the three bases C, T, and ANYTHING ELSE, you make leucine.
This is pretty good for leucine, because after CT you can have thymine or cytosine or guanine or adenine next, and it doesn't make any difference. You've still produced leucine. Leucine is very forgiving! TTA, or TTG will ALSO make leucine.
Look at histidine, though. Can you see that it's pickier? It's CAT or CAC - and that's it. If you screw one of those up, no histidine for you.
And stuff like methionine? You only have one chance to get it right. The codon for methionine is ATG - that's it. Let's say you have a SNP that causes you to make ATC instead. Now you've made isoleucine instead of methionine. Whoops!
That's a missense mutation: where the mutation causes you to produce the wrong amino acid.
The consequences of this could be dire or minimal (like all of this - sorry, guys). Amino acids hook together to make every protein in your body. If the protein is still functional with the alteration, it's all good. If the alteration changes the shape of the protein to make it harder to fit into receptors or carry molecules - the way hemoglobin carries oxygen and carbon dioxide, for example - or not fit into the proper receptors at all - or makes it unstable so that the protein is fragile - then you could be in for a world of trouble.
BACK to our regularly scheduled programme:
See the gene? VWF... VWF... Von Willebrand Factor.
Uh oh. That gene helps blood clot. It's important!
Note where it says HGVS; at the bottom, you'll see Pro2063Ser. That tells us that if you have this SNP, in the VWF gene, proline was replaced with serine.
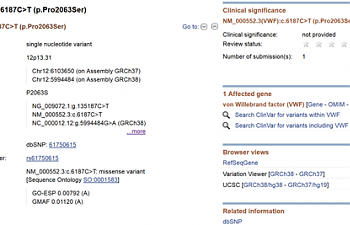
Click on the link for the SNP itself and...
And here we are once more. Click on ClinVar.
...wow. Not a single star's worth of evidence. One submission discussing the SNP, that says it affects VWF gene. Nothing we didn't already know. Aw, shucks. But don't give up yet.
There are two avenues to explore, yet.
To be Continued!
<---Previous in the series
<---------First in the series
There are very few SNPs that are the subject of so many studies as rs234706 (even through the worst brain fog, I will always remember that SNP!)
Let's take Rs61750615 as our example, this time. It's one of my rarest SNPs, occurring in less that one half of 1% of the population overall.
(Sounds super-duper rare, guys, but it's still one out of about 200 people, implying that there were maybe five at my old high school with this SNP.)
If you try to go to snpedia, this is what you'll see:
A page inviting you to please create a page for this SNP! They appear to have no information whatsoever.
Okay, let's try our tried-and-true method described in the previous post. Go to NCBI's page, drop-down menu, select SNP, and in the search box, type the rs#. This shows up:
Well, whaddaya know? We got a little something out of it this time.
If we look where it says 'functional consequence' the page says 'missense'. This is a genetics term to avoid saying 'nonsense', but it has the same connotation.
Each three bases in a row - like A, G, G, for example - is called a codon. And codons are called that because they each code for amino acids:
So, if you have the three bases C, T, and ANYTHING ELSE, you make leucine.
This is pretty good for leucine, because after CT you can have thymine or cytosine or guanine or adenine next, and it doesn't make any difference. You've still produced leucine. Leucine is very forgiving! TTA, or TTG will ALSO make leucine.
Look at histidine, though. Can you see that it's pickier? It's CAT or CAC - and that's it. If you screw one of those up, no histidine for you.
And stuff like methionine? You only have one chance to get it right. The codon for methionine is ATG - that's it. Let's say you have a SNP that causes you to make ATC instead. Now you've made isoleucine instead of methionine. Whoops!
That's a missense mutation: where the mutation causes you to produce the wrong amino acid.
The consequences of this could be dire or minimal (like all of this - sorry, guys). Amino acids hook together to make every protein in your body. If the protein is still functional with the alteration, it's all good. If the alteration changes the shape of the protein to make it harder to fit into receptors or carry molecules - the way hemoglobin carries oxygen and carbon dioxide, for example - or not fit into the proper receptors at all - or makes it unstable so that the protein is fragile - then you could be in for a world of trouble.
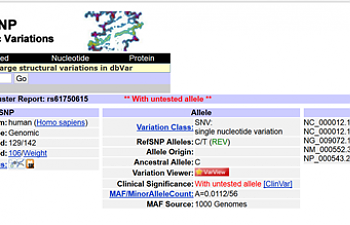
BACK to our regularly scheduled programme:
See the gene? VWF... VWF... Von Willebrand Factor.
Uh oh. That gene helps blood clot. It's important!
Note where it says HGVS; at the bottom, you'll see Pro2063Ser. That tells us that if you have this SNP, in the VWF gene, proline was replaced with serine.
Click on the link for the SNP itself and...
And here we are once more. Click on ClinVar.
...wow. Not a single star's worth of evidence. One submission discussing the SNP, that says it affects VWF gene. Nothing we didn't already know. Aw, shucks. But don't give up yet.
There are two avenues to explore, yet.
- Go back to pubmed and cut-and-paste that rs# into the searchbox. This will catch any articles that mention the SNP.
Pubmed turns up nothing. 
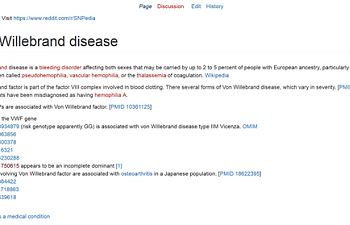
- Go back to snpedia and search for Von Willebrand. Maybe it mentions the SNP, even if that SNP doesn't have its own page.
...And snpedia delivers.
To be Continued!
<---Previous in the series
<---------First in the series